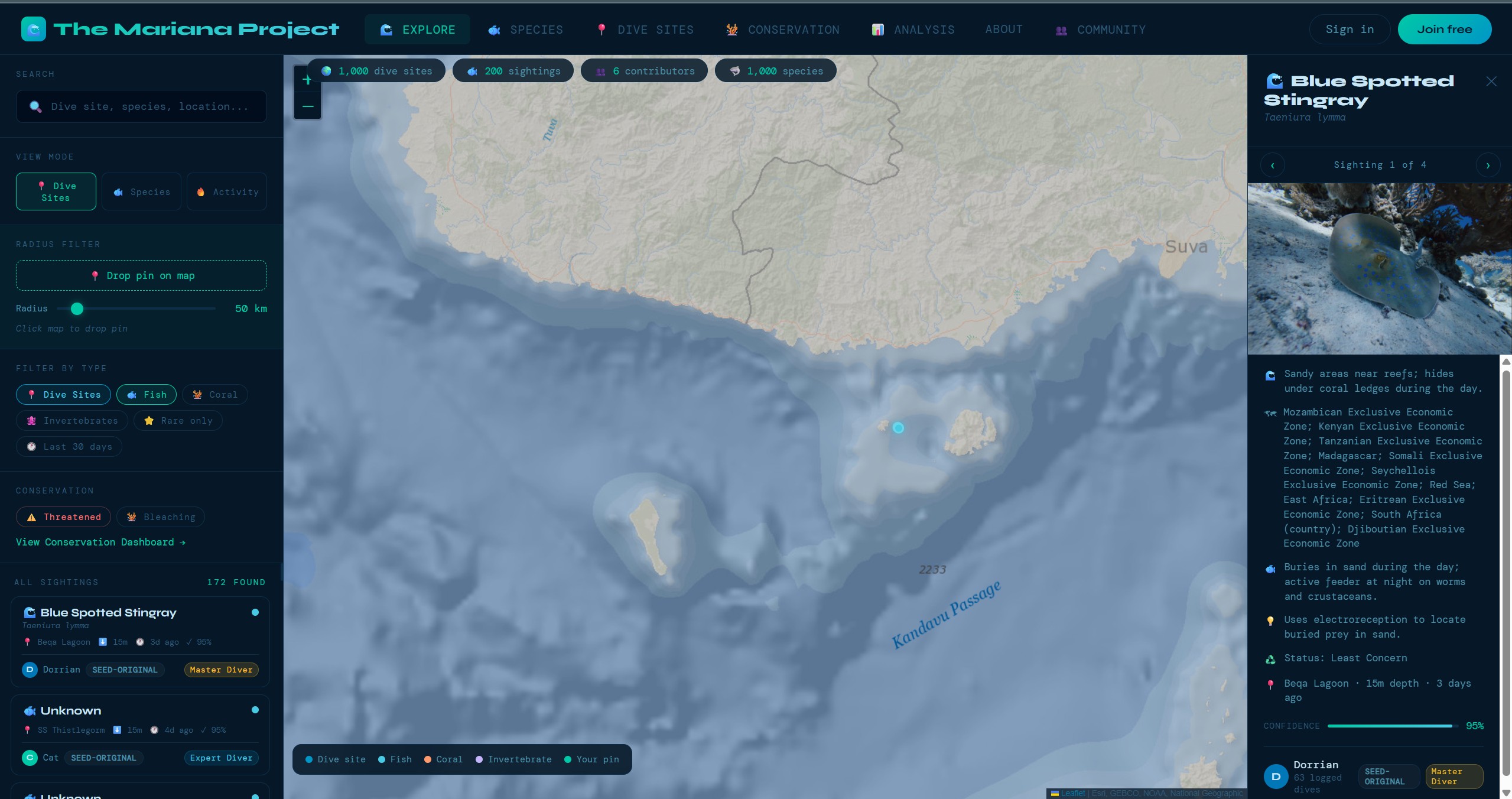
"AI-assisted marine biodiversity platform for divers, communities, and researchers"
A citizen science platform designed to turn diver observations into structured marine biodiversity records. Combines AI-assisted species identification, interactive mapping, community verification, and standards-based export — built solo with a strong focus on data quality and real-world usability.

Marine biodiversity data exists across research databases, government sources, local projects, and personal dive logs — but it is fragmented, inconsistently structured, and often inaccessible to the wider public. That makes it harder than it should be to build a useful picture of what is being seen, where, and by whom.
At the same time, millions of recreational divers spend time underwater observing species, photographing marine life, and logging dives. That is potentially valuable ecological signal, but most of it never becomes structured, reusable data because the contribution process is too technical or too time-consuming.
Existing citizen science tools often solve only part of the problem: they may be easy to use but weak on validation, or rigorous but too specialist for everyday contributors. The opportunity here was to make it easy for divers to contribute while still producing data researchers could meaningfully work with.
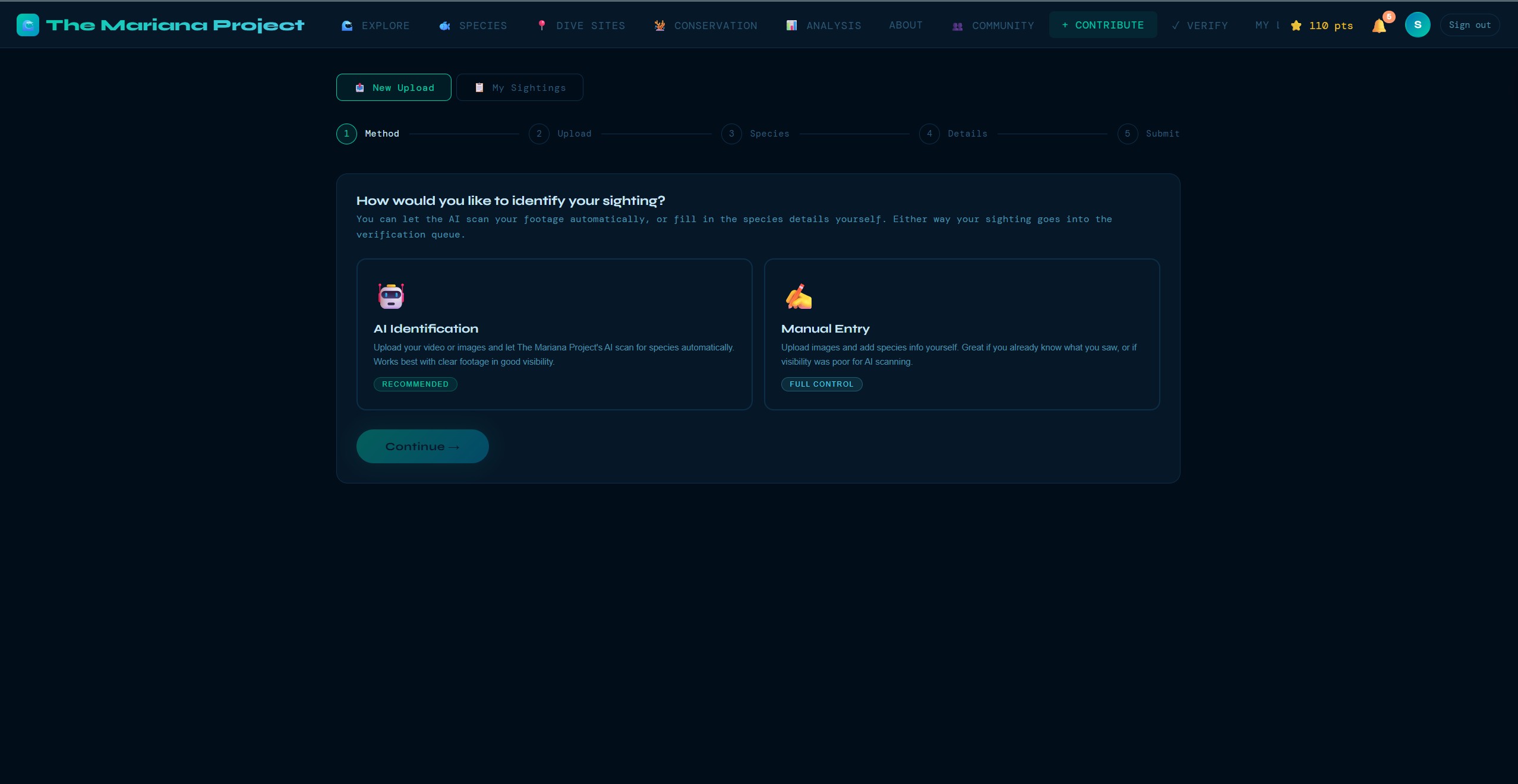
The Mariana Project is designed to make contributing biodiversity data feel closer to logging a dive than filling in a research form. Divers submit sightings with a photo, a location, and the relevant details; the platform then layers in AI-assisted identification, community review, validation checks, and standards-based export for downstream research use.
The aim is not to pretend every observation is instantly research grade. The aim is to create a workflow that takes casual observations and improves their usefulness through structure, validation, and transparent review — so more ocean activity becomes usable signal over time.
Every technology choice in The Mariana Project was deliberate. The goal was maximum functionality with minimum operational complexity — a solo developer needs tools that punch above their weight.
No framework overhead. Faster load times, simpler deployment, full control. For a content-heavy platform with interactive maps, vanilla JS with modern APIs (Fetch, Intersection Observer) handles everything without a build step.
Leaflet is lightweight and plugin-rich. ESRI's Ocean Basemap is purpose-built for marine applications — bathymetric contours, ocean features, and reef systems rendered beautifully at every zoom level.
A full backend-as-a-service with the power of PostgreSQL. Row-Level Security policies enforce data access at the database level. Auth, storage, and real-time subscriptions — all from one platform, all with a generous free tier.
Claude's vision capabilities power the species identification pipeline. A diver uploads a photo; Claude analyses it against taxonomic knowledge and returns a structured identification with confidence scoring and a common-name explanation.
The World Register of Marine Species and Ocean Biodiversity Information System provide taxonomic authority. Every species in the database is validated against WoRMS for correct nomenclature, and OBIS provides distribution data for plausibility checks.
Static site hosting with edge CDN, automatic deploys from Git, and built-in form handling. Zero server management, instant rollbacks, and free SSL — ideal for a frontend-heavy app backed by Supabase.
Upload a photo or frame from video footage and Claude analyses the image, identifies the species with a confidence level, provides the scientific name (validated against WoRMS), common name, and conservation status. Designed to work even with partial or blurry underwater photos.
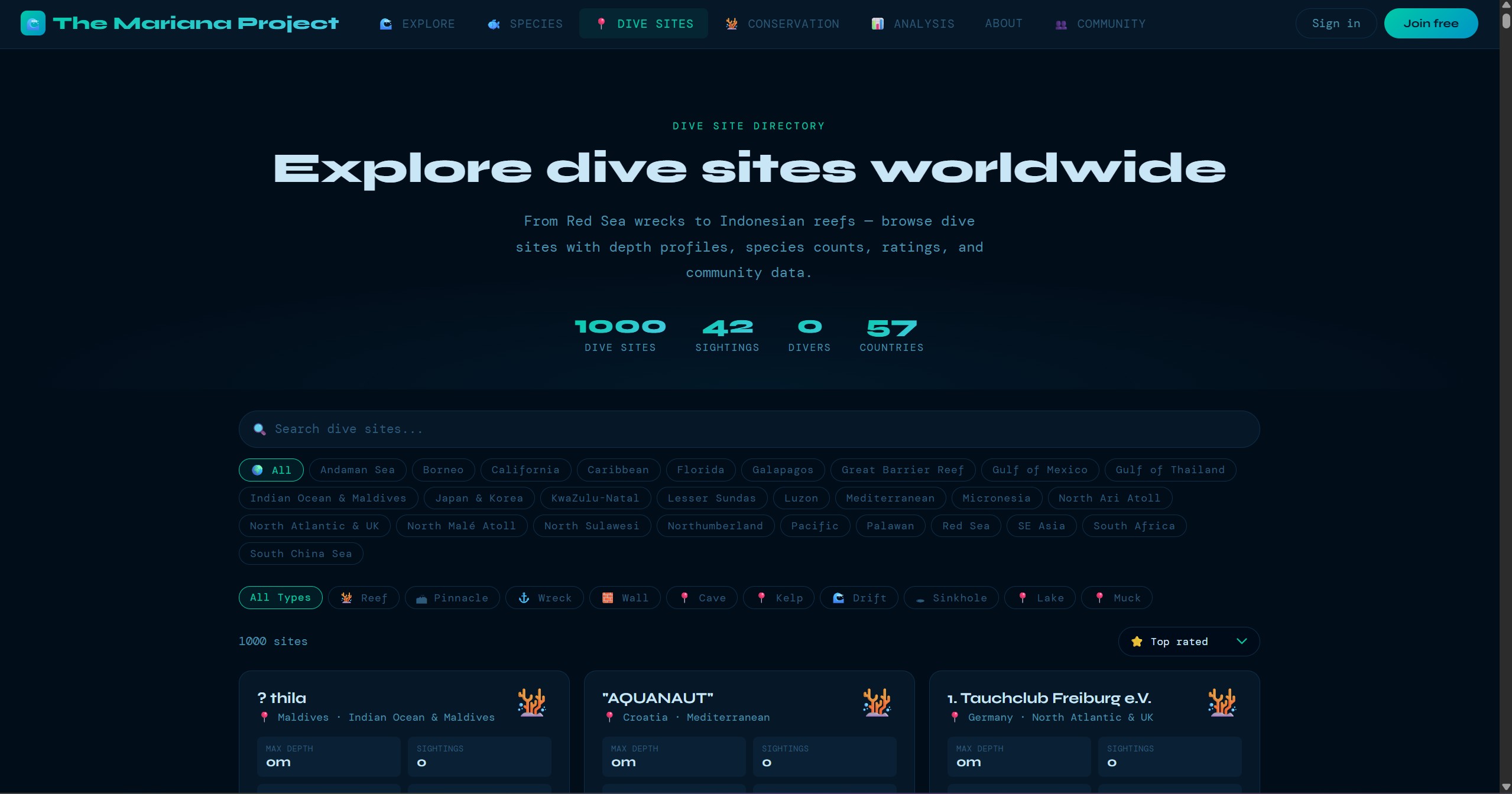
1,850+ dive sites plotted on an ESRI Ocean Basemap via Leaflet.js. Sites are filterable by region, clustered for performance at low zoom levels, and each site links to detailed information including conditions, species lists, and community sightings.
A full dive logging system — depth, duration, conditions, buddy, equipment, notes — with species life lists that build automatically from verified sightings. Every dive becomes both a personal record and a data contribution.
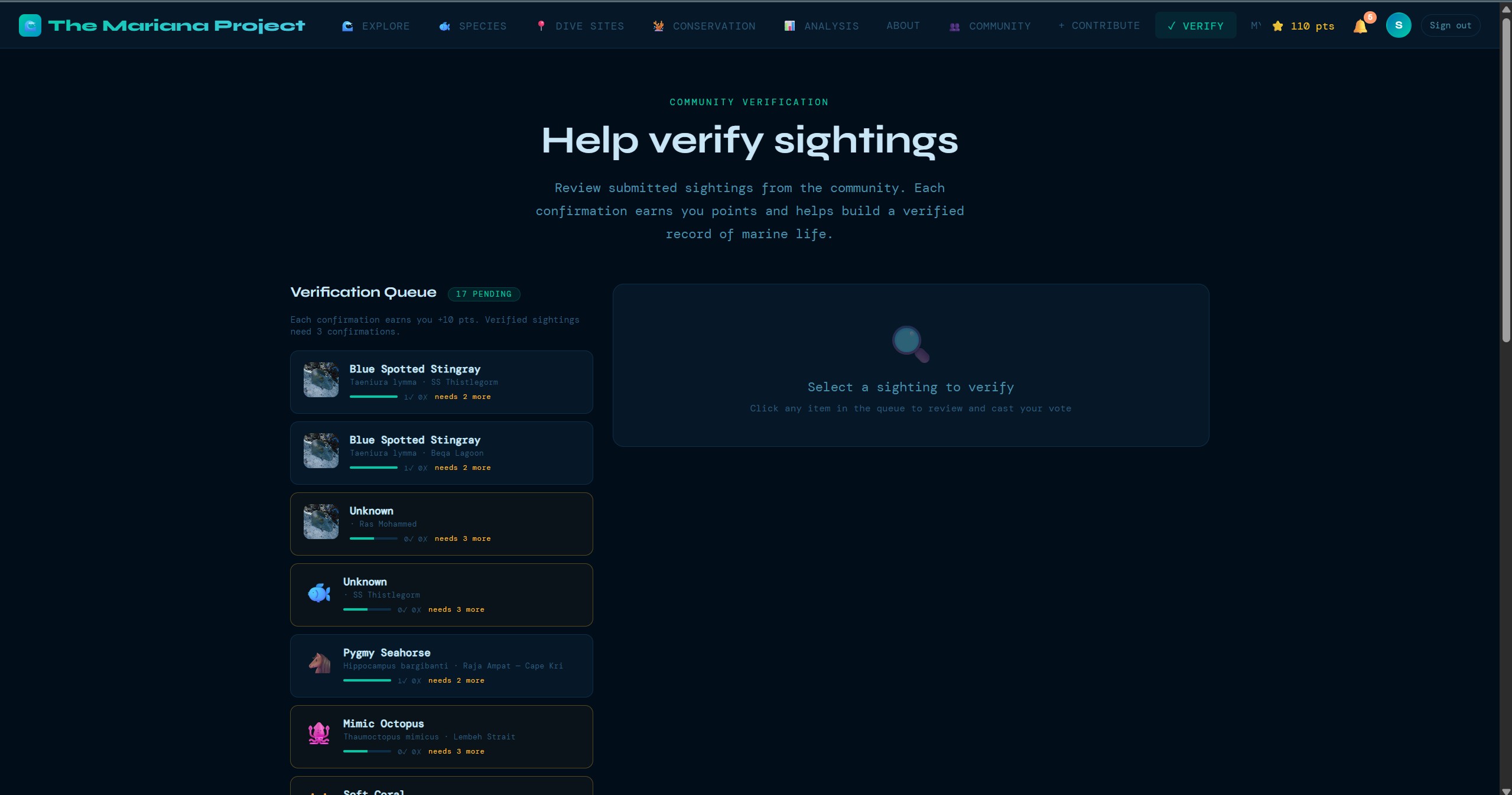
A multi-stage verification system where experienced community members review and validate sightings. Consensus-based scoring builds data confidence over time, with disagreements flagged for expert review.
Every sighting passes through a 26-point validation pipeline checking taxonomic validity, geographic plausibility, temporal consistency, duplicate detection, image quality assessment, and more — before it enters the verified dataset.
Verified data exports in Darwin Core standard — the international format used by GBIF and OBIS. This makes The Mariana Project data directly usable by marine researchers, conservation organisations, and policy makers worldwide.
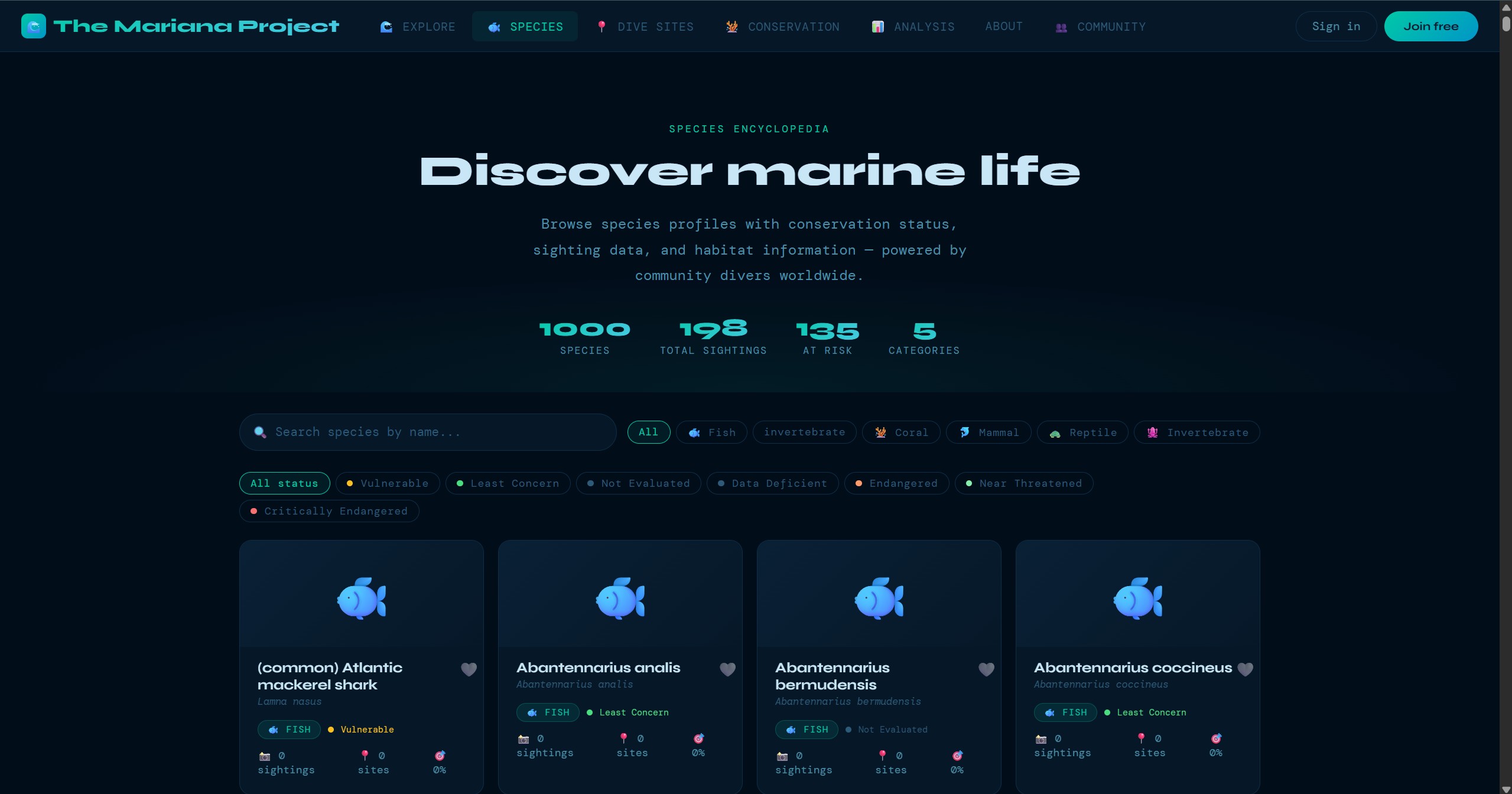
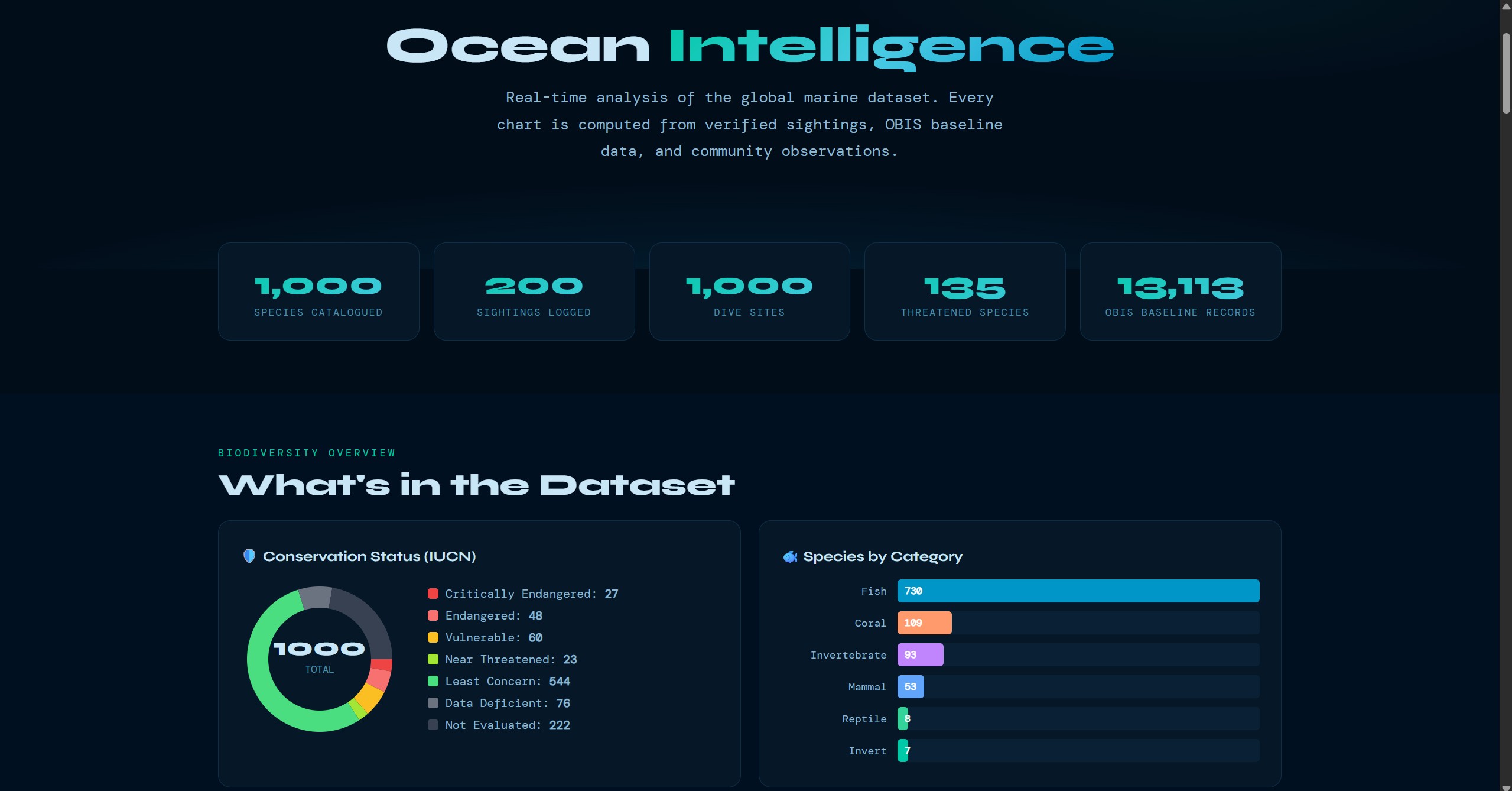
These screens show the product beyond the headline map view: guided submission, dive-site exploration, species browsing, community verification, and the data views that support conservation analysis.
Challenge: WoRMS and OBIS APIs return data in different formats with varying completeness. Some species had multiple accepted names, others had missing distribution records, and rate limits made bulk imports slow.
Solution: Built a custom ETL pipeline in Python that normalises records from both APIs into a consistent schema, deduplicates on AphiaID (the WoRMS unique identifier), handles rate-limiting with exponential backoff, and logs every transformation for audit. The pipeline processes species in batches and validates each record before insert.
Challenge: Underwater photography is notoriously difficult — low light, particulate matter, moving subjects, and colour distortion from depth. A naive image classification approach would fail on most real-world dive photos.
Solution: Engineered detailed prompts for Claude that account for underwater conditions. The system asks for multiple possible identifications ranked by confidence, flags when a photo is too degraded for reliable ID, and cross-references against known species distributions for the dive site's geographic region to filter implausible matches.
Challenge: Research-grade data demands rigorous validation, but overly strict checks would reject legitimate sightings and discourage contributors. Finding the balance between data quality and contributor experience was critical.
Solution: Implemented a tiered validation system: hard checks (invalid coordinates, non-existent species) block submission; soft checks (unusual depth range, uncommon species for region) flag for review but allow through. Each check produces a structured log entry, making the pipeline fully auditable and tuneable over time.
Beyond using Claude's general vision capabilities, The Mariana Project includes a custom AI training pipeline designed to fine-tune models specifically for marine species identification. Using QLoRA (Quantized Low-Rank Adaptation), the pipeline allows efficient fine-tuning on consumer hardware — making it possible to build specialised identification models without cloud GPU costs.
The training data comes from the platform's own verified sightings, creating a feedback loop: as more divers contribute and the community verifies sightings, the training dataset grows, and the AI improves. This self-reinforcing cycle is core to The Mariana Project's long-term vision.
The Mariana Project is under active development with a substantial roadmap. Near-term priorities include community groups for dive clubs and research teams, stronger contribution loops, and deeper integration pathways with GBIF and OBIS so verified data can move more cleanly into the wider biodiversity ecosystem.
Longer term, the platform expands into a mobile companion experience for offline dive logging, better alerting for unusual sightings, and a more open data layer for researchers and conservation partners who want to explore or reuse the dataset programmatically.